Research Interests
Viruses infect their respective hosts efficiently through a regulated process of recognizing highly specific receptors and subsequently transferring genomic material across cell membrane barriers. To understand the underlying mechanisms that control virus infection, it is important to characterize viral events in a biologically relevant manner. My laboratory focuses on a correlated approach to deciphering the processes of virus assembly and infection via a combination of biochemistry, molecular biology, biophysics and structural biology (electron cryo-microscopy and three-dimensional image reconstruction methods). The main project of the laboratory involves understanding assembly and host-recognition mechanisms in dsDNA-containing bacteriophage (family Podoviridae).
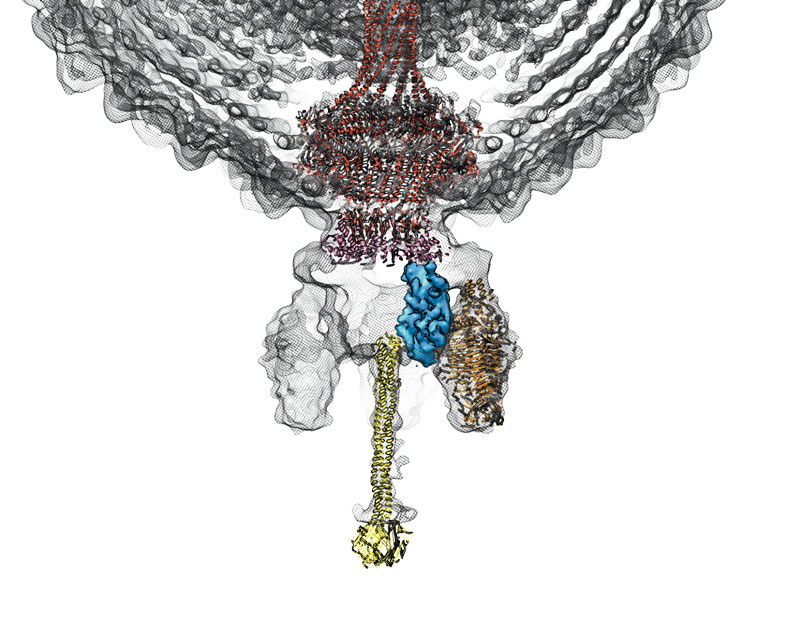
Colored in yellow is the tail needle (gp9)
I have been able to study Sf6 infection in the context of its host, Shigella flexneri, through classic phage analyses and have identified two secondary receptors that are crucial to Sf6 infection (S. flexneri outer membrane proteins A & C, "Omps A & C"). We are studying the interactions between Sf6 and S. flexneri to determine regions in the phage tail and in the Omps that are critical for proper infection. We are extending our understanding of Sf6 infection dynamics since this phage mimics infection in vitro by binding and delivering its genome into lipid vesicles comprised of host membranes and Omps A and C. This provides an ideal system for using cryo-electron tomography to visualize intermediates that arise during Sf6 infection, as the lipid vesicles are considerably thinner (~100 nm thick) compared to whole Shigella cells (~2000 nm thick), which are currently outside the limits of effective cryo-tomography. This makes Sf6 one of a few model systems to correlate structural transitions that arise during host cell recognition and resulting genomic transfer. This, combined with decades of groundwork on bacteriophage genetic manipulation and biochemical/biophysical characterization, allows us to study the process of Sf6 infection from several different angles and will provide insight into the generalized mechanisms by which all viruses recognize, attach, and infect hosts.